Querying Component
PATIKAweb provides an advanced, graph-based querying facility for retrieving the data of user's interest from the PATIKA database. Alternately, the user may load its model locally (using either one of PATIKA or BioPAX level 2 formats), and perform a query on this model. In either case, we assume the model (i.e. a pathway knowledgebase stored in a database or loaded up into memory) is made up of integrated pathways as opposed to separate independent set of pathways as shown below.

Querying component of PATIKA both supports SQL-like queries and an array of graph-theoretic queries for finding feedback loops, positive/negative paths, common targets and regulators, shortest paths, or "interesting subgraphs" based on user's genes of interest. Once retrieved from the database, the query results may be merged to the user's current model and highlighted to provide an incremental, user-friendly retrieval and analysis interface. Constructed models can be saved in a native XML-based format (".pmdl"), exported to standard formats such as BioPAX level 1 and SBML, or converted to static images.
The query interface of PATIKAweb has been implemented as an applet. In the following sections you will find sample queries presented with screenshots from this applet.
Query Types
Field Queries
The simplest query type that can be performed in PATIKAweb through the query applet is the field query. The database may be queried using many types of fields including name, PATIKA ID, description, version and GO terms.
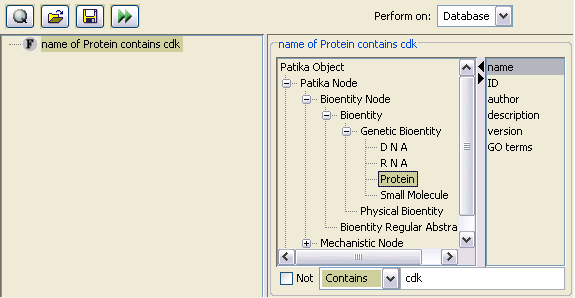
You may specify the PATIKA object type you'd like to search for, using the PATIKA object tree. Please refer to the ontology section for a better understanding of the PATIKA object tree.
Other types of queries use the results of field queries as input (e.g. source and target node sets for a shortest path query, or node set whose neighbors are to be found for a neighborhood query).
Combining Queries through Logical Operators ("AND" and "OR")
It is possible to combine two or more queries with AND and OR logical operators. The new construct is also a query, that may recursively be used in other AND and OR queries, or in other types of queries.

States, Sources and Products of Bioentities
All states (mechanistic level nodes) of a specified bioentity (bioentity level nodes) may be queried using "States of a Bioentity" query.
Source (product) bioentities of a specified bioentity (e.g. source DNA of a protein) may be queried through "Sources (Products) of a Bioentity" query.
Graph-Theoretic Queries
Graph algorithms such as depth/breadth-first traversals and shortest paths are used to compute feedback loops, positive/negative paths, common targets and regulators, or "interesting subgraphs" based on user's interest of genes. Please refer to the User's Guide for details.
Query Results
After the execution of a query initiated from the Query Dialog finishes, the returning result (i.e. pathway model) is summarized by the Query Result Dialog.
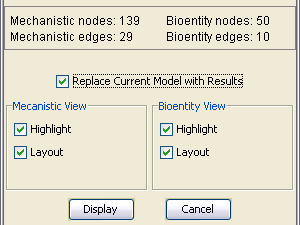
A number of statistics about the result is displayed in this dialog:
The user may opt to highlight the objects comprising the query result in each of the views.

In addition, the user decides whether the resulting views are to be laid out before displaying.
A Querying Scenario
Following is a sample session in which subsequent queries and complexity management operations are performed to form a model that might be of use to a PATIKAweb user.
Suppose the user is studying the effects of FAS Ligand on apoptosis. One good way to start is by searching for the relations between FAS Ligand and the Caspase complexes in the cell.
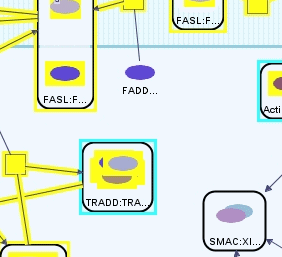
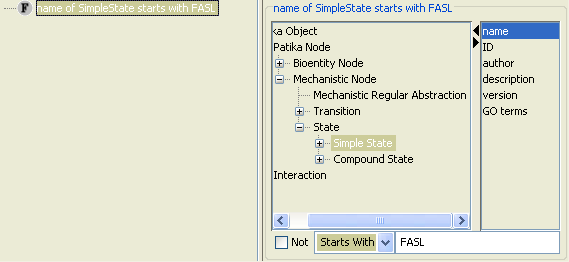
In order to find out the states of FAS Ligand in the cell, we perform the query in Figure 6, where we ask for simple states whose names start with FASL.
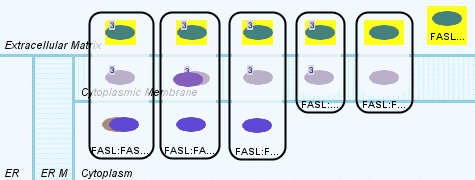
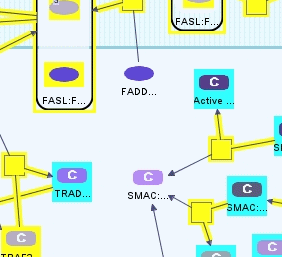
We see 6 states highlighted in the result of the query (Figure 7). One is the free extracellular FAS Ligand, and remaining ones are members of several complexes spanning the cytoplasmic membrane.
And we may check how many Caspase complexes we have in the database, which are not a precursor or a pro-caspase (Figure 8).
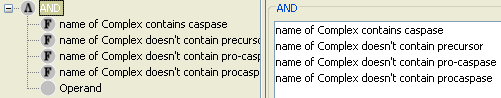
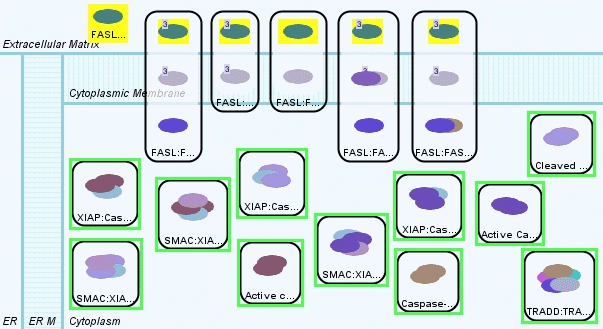
Caspase query returns a total of 11 complex molecules, which are all in cytoplasm (Figure 9). Now we know that the database contents that we want to start from and we want to reach to. The most popular query for finding relatively short paths between source and target molecules is the Shortest Path Query. We may use the previous FAS Ligand and Caspase field queries as the source and target fields of the shortest path query (Figure 10).
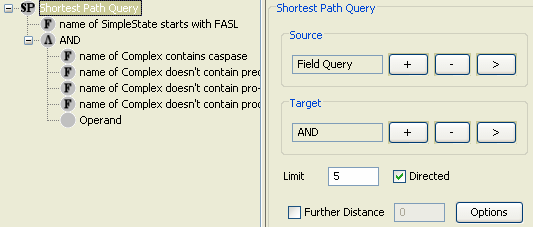
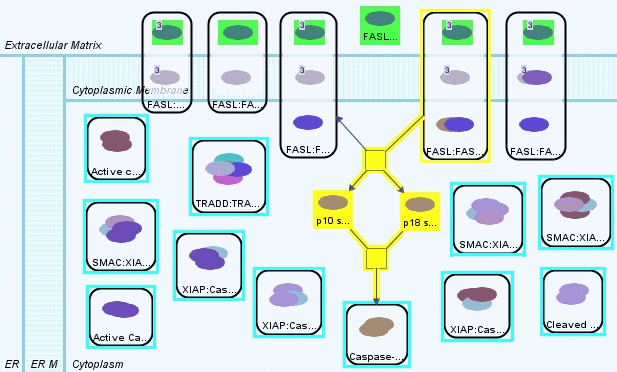
Result of the shortest path query retrieves paths of length 4 (Figure 11). These are paths involving the FAS Ligand complex on the cytoplasmic membrane and the Caspase-8 dimer in cytoplasm. This picture might be very helpful but it still has many missing relations.
There are several ways to obtain a more complete picture. First alternative is to use the shortest path query with the Further Distance parameter. Figure 12 shows the same query with further distance set to 8. Since the shortest path length is 4, this query would bring us the paths from source to target nodes of length at most 12. Figure 13 shows the resulting model.
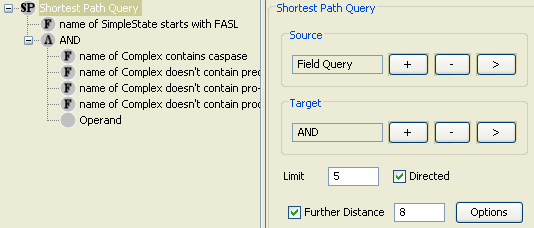
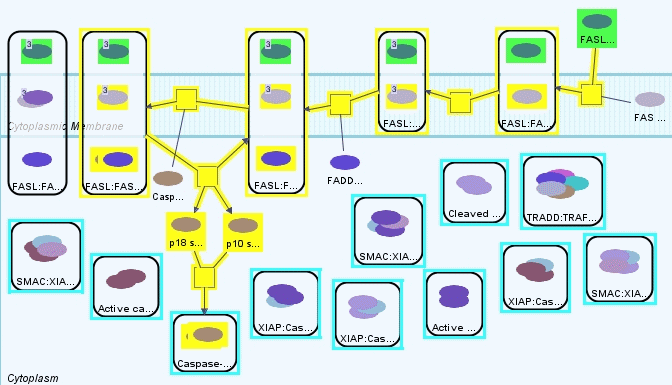
Another way of doing the same query is to use a Paths-of-Interest (PoI) query with limit 12. Since this query will bring all paths of length at most 12, between source and target sets, the result will be identical to the previous shortest path query with further distance 8. Thus PoI query is simply a more convenient way of querying paths when we have a good estimation of the length of the shortest path.
When finding paths between source and target sets is not sufficient, the user has the option to do a Graph-of-Interest (GoI) query. GoI query aims at completing the missing links (and molecules on these links) among a set of molecules of interest that is no longer than a specified limit. So a minimal graph including the specified objects of interest can be constructed through this query. Figure 14 shows a directed GoI query with limit 5, where the previous source and target sets are joined into an OR query as molecules of our interest.Since the GoI query finds all paths between a number of seed nodes (not from a specified source to specified target), the result contains more paths, not necessarily depicting a direction in the information flow. In the resulting model (Figure 15) we see that there are two isolated components. First one contains the previous FAS Ligand path we have found. We see that an additional Caspase complex is connected; however, the graph does not imply that this new Caspase complex has been involved in the FAS Ligand signaling process. Second component contains all other Caspase complexes. Notice that only two Caspase complexes have a relation with FAS Ligand signaling process in the database (at least within the distance we have specified); the user may choose to concentrate on these for further analysis.